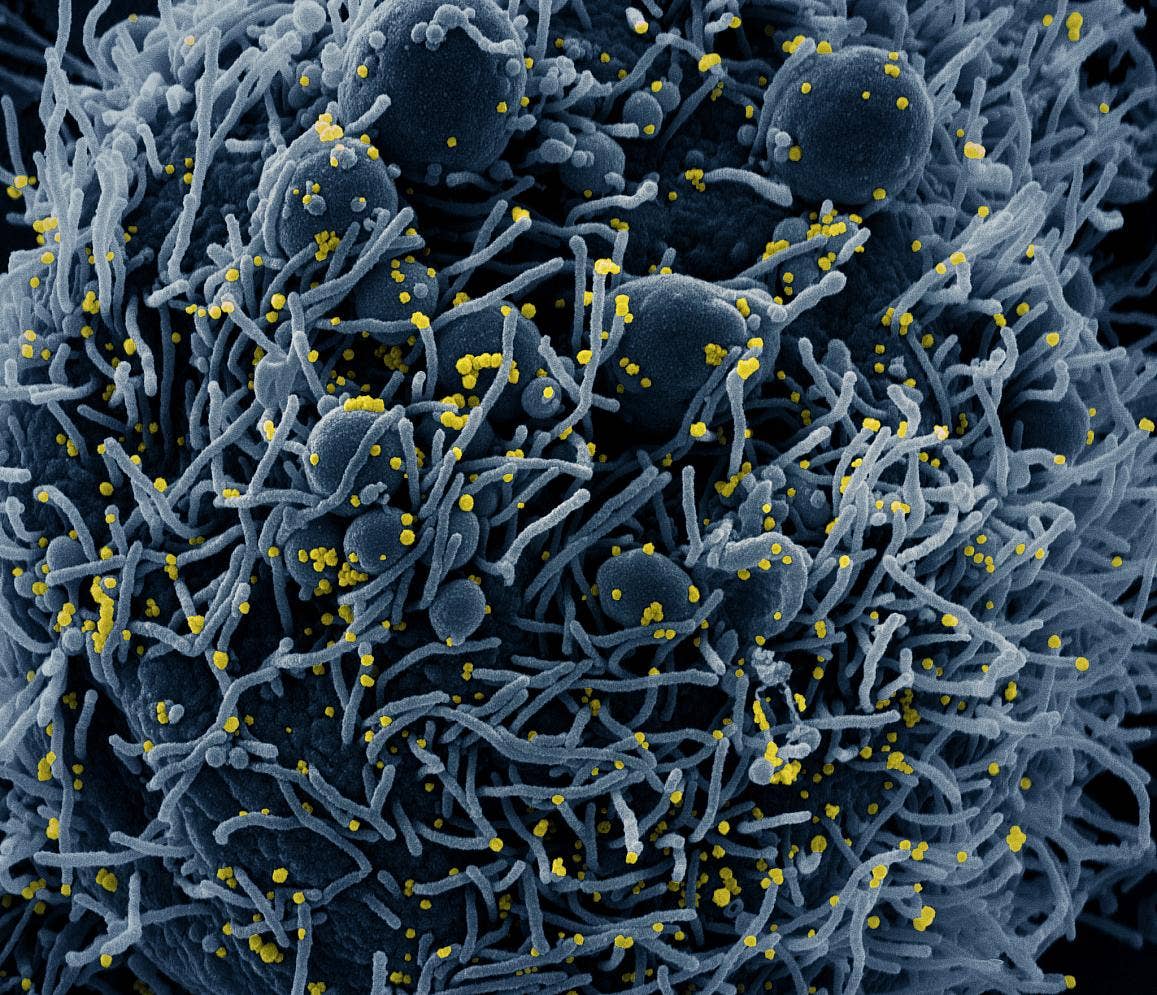
This claims specifically there is no HIV and again I would have to read carefully to be sure.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7033698/

Infection from an emerging pathogenic coronavirus was first reported in December 2019 in China. It has now affected over 42,000 people and caused over 1,000 deaths in 25 countries (https://2019ncov.Chinacdc.Cn/2019-Ncov). The complete genome of this new virus was quickly sequenced and made public on January 12, only about 2 weeks after the disease was first observed [4]. It was named as 2019-nCoV the following day by the World Health Organization (WHO). Phylogenetic analysis shows that 2019-nCoV is a new member of coronaviruses that infect humans. It is genetically homogenous but distinct from coronaviruses that cause SARS and MERS [5,6]. However, it shares a high level of genetic similarity (96.3%) with a bat coronavirus RaTG13 which was obtained from bat in Yunnan in 2013, suggesting that RaTG13-like viruses are most likely the reservoir, but not the immediate sources of the current 2019-nCoV viruses [7].
Lack of the definite origin of 2019-nCoV has led to speculation that 2019-nCoV might be derived from genetic manipulation or even for the purpose of use as a bioweapon. This notion has been fully debunked in the media. A recent informally presented report, however, showed that 2019-nCoV had four insertions in the spike glycoprotein gene that is critical for the virus to enter the target cells when compared to other coronaviruses [8]. It was claimed that these inserts were either identical or similar to the motifs in the highly variable (V) regions (V1, V4 and V5) in the envelope glycoprotein or in the Gag protein of some unique HIV-1 strains from three different countries (Thailand, Kenya and India). Together with the structure modelling analysis, the authors speculated that these motif insertions sharing similarity with HIV-1 proteins could provide an enhanced affinity towards host cell receptors and increase the range of host cells of 2019-nCoV. This study implies that 2019-nCoV might be generated by gaining gene fragments from the HIV-1 genome.
Current report conducted careful examination of the sequences of 2019-nCoV, other CoV viruses and HIV-1 as well as GenBank database. Our results demonstrated no evidence that the sequences of these four inserts are HIV-1 specific or the 2019-nCoV viruses obtain these insertions from HIV-1. First, the results of blast search of these motifs against GenBank shows that the top 100 identical or highly homologous hits are all from host genes of mammalian, insects, bacterial and others. There are only a few hits on coronaviruses, but none of them are HIV-1 related. Blast against viral sequence database also showed these insertion sequences widely exist in all kinds of viruses from bacteriophage, influenza, to giant eukaryotic viruses (Table 1). More hits were found for coronaviruses and a few also hit on HIV-1 sequences than the search against the entire database (Table 1). However, while the 100% match between the insertion 1 and 2 sequences and the HIV sequences were found in 19 entries, the matches between the insertion 3 and 4 sequences and HIV-1 sequences were rather poor (from 42% to 88%). Moreover, the insertion 4 sequence ambiguously hit multiple different genes (gag, pol and env) in the HIV-1 genome, suggesting that similarities (as low as 42%) between them are too low to be reliable. Search these four insertion sequences against HIV-1 Sequence Database (https://www.hiv.lanl.gov/components/sequence/HIV/search/search.html) yielded similar results. Sequences that completely match the insertion 3 and 4 sequences were not found in any HIV-1 sequences. This clearly shows that these insertioin sequences are widely present in living organisms including viruses, but not HIV-1 specific. All these regions in HIV-1 envelope glycoprotein are highly variable with many large insertions and deletions, indicating that they are not essential for biological functions of HIV-1 envelope glycoprotein. The detection of completely matched sequences of 1 and 2 insertions in only a few HIV-1 strains demonstrated that four insertions are very rare or not present among tens of thousands of natural HIV-1 sequences. This also explains why four insertion homolog sequences could only be independently found in different HIV-1 genomes [8]. Because of their poor identities to and rareness in the HIV-1 sequences, HIV-1 could not be the source for those insertion sequences in the 2019-nCoV genome.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7033698/
Infection from an emerging pathogenic coronavirus was first reported in December 2019 in China. It has now affected over 42,000 people and caused over 1,000 deaths in 25 countries (https://2019ncov.Chinacdc.Cn/2019-Ncov). The complete genome of this new virus was quickly sequenced and made public on January 12, only about 2 weeks after the disease was first observed [4]. It was named as 2019-nCoV the following day by the World Health Organization (WHO). Phylogenetic analysis shows that 2019-nCoV is a new member of coronaviruses that infect humans. It is genetically homogenous but distinct from coronaviruses that cause SARS and MERS [5,6]. However, it shares a high level of genetic similarity (96.3%) with a bat coronavirus RaTG13 which was obtained from bat in Yunnan in 2013, suggesting that RaTG13-like viruses are most likely the reservoir, but not the immediate sources of the current 2019-nCoV viruses [7].
Lack of the definite origin of 2019-nCoV has led to speculation that 2019-nCoV might be derived from genetic manipulation or even for the purpose of use as a bioweapon. This notion has been fully debunked in the media. A recent informally presented report, however, showed that 2019-nCoV had four insertions in the spike glycoprotein gene that is critical for the virus to enter the target cells when compared to other coronaviruses [8]. It was claimed that these inserts were either identical or similar to the motifs in the highly variable (V) regions (V1, V4 and V5) in the envelope glycoprotein or in the Gag protein of some unique HIV-1 strains from three different countries (Thailand, Kenya and India). Together with the structure modelling analysis, the authors speculated that these motif insertions sharing similarity with HIV-1 proteins could provide an enhanced affinity towards host cell receptors and increase the range of host cells of 2019-nCoV. This study implies that 2019-nCoV might be generated by gaining gene fragments from the HIV-1 genome.
Current report conducted careful examination of the sequences of 2019-nCoV, other CoV viruses and HIV-1 as well as GenBank database. Our results demonstrated no evidence that the sequences of these four inserts are HIV-1 specific or the 2019-nCoV viruses obtain these insertions from HIV-1. First, the results of blast search of these motifs against GenBank shows that the top 100 identical or highly homologous hits are all from host genes of mammalian, insects, bacterial and others. There are only a few hits on coronaviruses, but none of them are HIV-1 related. Blast against viral sequence database also showed these insertion sequences widely exist in all kinds of viruses from bacteriophage, influenza, to giant eukaryotic viruses (Table 1). More hits were found for coronaviruses and a few also hit on HIV-1 sequences than the search against the entire database (Table 1). However, while the 100% match between the insertion 1 and 2 sequences and the HIV sequences were found in 19 entries, the matches between the insertion 3 and 4 sequences and HIV-1 sequences were rather poor (from 42% to 88%). Moreover, the insertion 4 sequence ambiguously hit multiple different genes (gag, pol and env) in the HIV-1 genome, suggesting that similarities (as low as 42%) between them are too low to be reliable. Search these four insertion sequences against HIV-1 Sequence Database (https://www.hiv.lanl.gov/components/sequence/HIV/search/search.html) yielded similar results. Sequences that completely match the insertion 3 and 4 sequences were not found in any HIV-1 sequences. This clearly shows that these insertioin sequences are widely present in living organisms including viruses, but not HIV-1 specific. All these regions in HIV-1 envelope glycoprotein are highly variable with many large insertions and deletions, indicating that they are not essential for biological functions of HIV-1 envelope glycoprotein. The detection of completely matched sequences of 1 and 2 insertions in only a few HIV-1 strains demonstrated that four insertions are very rare or not present among tens of thousands of natural HIV-1 sequences. This also explains why four insertion homolog sequences could only be independently found in different HIV-1 genomes [8]. Because of their poor identities to and rareness in the HIV-1 sequences, HIV-1 could not be the source for those insertion sequences in the 2019-nCoV genome.